New Framework Quantifies Protein Embedding Reliability
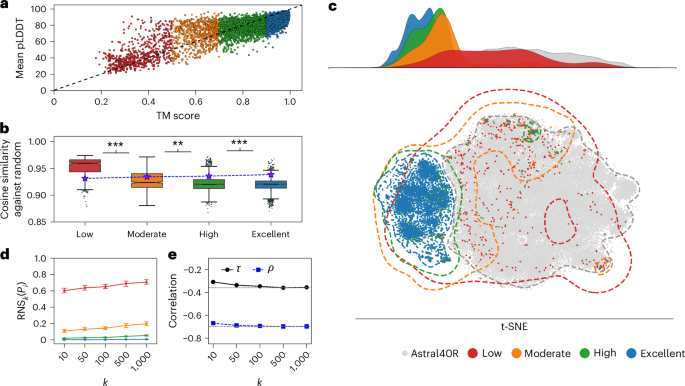
A recent study has introduced a novel scoring framework aimed at assessing the reliability of biomolecular embeddings used in protein language models. This framework quantifies representation uncertainty by measuring the fraction of biologically meaningless sequences present among a protein's nearest neighbors in the latent space. The findings underscore a critical flaw in utilizing embeddings without validating their biological relevance, revealing that low-quality embeddings can closely resemble randomly generated sequences, potentially leading to misleading biological insights.
The implications of this research are significant for the field of genome informatics and machine learning applications in biology. By advocating for routine embedding evaluations prior to downstream applications, the study enhances the reliability of predictions made using protein language models. This approach is not only poised to improve the accuracy of protein function predictions and mutation effects but also sets a precedent for the evaluation of language models across various scientific disciplines, ensuring that machine learning applications adhere to stringent standards of reliability.